
The Disease Detective
UW researcher Tony Goldberg is on the hunt for deadly viruses.
If you are Tony Goldberg, the unknown agents of disease are right under your nose. Sometimes they are even in your nose.
For Goldberg, a professor of pathobiological sciences in UW–Madison’s School of Veterinary Medicine, the ever-lurking emissaries of infectious disease — viruses, bacteria, fungi, parasites — are the epidemiological Easter eggs that nature sprinkles on the landscapes of the world. Hiding in soil, water, plants, insects, and, frequently, other animals are pathogens that seemingly spring from nowhere to sow misery, death, and economic ruin.
Only a fraction of the world’s pathogens are known. More often than not, these organisms pose little or no threat to people, although they can cause disease in other mammals, fish, or plants. On occasion, though, they can jump to a new host species, including humans, whose immune defenses may not be prepared to ward off the invading microorganism.
Goldberg and many other scientists seek to identify pathogens before they emerge to threaten human and animal health. “We want to understand how pathogens in real ecosystems are transmitted,” explains Goldberg, who also serves as associate director for research at the UW’s Global Health Institute. “Our goal is to catch emerging diseases in the early stages so that we can contain them.”
The quintessential example in our lifetime — and the one that steered Goldberg from studying chimpanzees and other primates toward studying emerging diseases in animals — is HIV, the virus that causes AIDS. An estimated 70 million people have been infected with HIV, and more than 35 million have died from the disease. Scientists now believe AIDS originated from simian immunodeficiency virus in chimpanzees in the forests of Central Africa and took root in humans possibly as long ago as the 1850s.
More benign, perhaps, but rather creepy, is the new species of tick Goldberg found lodged in his nose after returning from fieldwork in Uganda’s Kibale National Park in 2013. The stowaway belonged to the genus Amblyomma and is a common parasite of chimpanzees. It makes a dash for the nose, scientists speculate, to avoid being groomed off. “Amblyomma ticks are known disease carriers, so this could be an underappreciated, indirect, and somewhat weird way in which people and chimps share pathogens,” Goldberg told an interviewer at the time.
Goldberg’s occupational-hazard-turned-scientific-discovery is just one in an astonishing series of disease-hunting adventures for the scientist whose boyish looks and good humor belie decades of tramping the world to track down pathogens. Goldberg and his team have worked for years in the lush forests of Uganda to seek out new viruses in monkeys and apes, animals that can be both the sources of new human diseases and also susceptible to human or livestock diseases that threaten their own health and conservation. Kibale National Park boasts an exceptional diversity of primates, many of which routinely come into contact with the people living in and around the park.
Closer to home, Goldberg spent a decade studying the spread of mosquito-borne West Nile virus in the suburbs of Chicago, where in 2012 he helped identify the American robin as a major player in the U.S. epidemic. West Nile is an opportunistic virus that has spread to at least 46 states and the District of Columbia since it was first documented in the United States in 1999. More recently, Goldberg has begun to plumb Wisconsin’s lakes and streams in search of pathogens that may be spreading in fish. A recent discovery by his group identified a new virus associated with largemouth bass dying off in large numbers in northern Wisconsin.
The task at hand for Goldberg and other disease hunters is extraordinarily difficult because, for starters, they often have no idea what they are looking for.
“How do you find a disease you don’t know anything about? We see a lot of cases in wildlife or people where we suspect an infection but we can’t quite figure out what it is,” he says. “Traditional methods of testing involve knowing something about what you’re looking for. You either use an antibody that detects a specific pathogen protein or a molecular test that targets the specific DNA sequence of a pathogen.”
Technology has advanced to the stage where scientists can randomly sequence millions of molecules of DNA in a sample and use powerful computing techniques to compare the resulting snippets of genetic material to known pathogens. When scientists get a promising “hit,” they can follow up with more targeted tests. “This is how you do virus hunting these days,” says Goldberg, noting the “deep sequencing” technique is used in his lab to ferret out viruses that affect a diversity of animals, from baboons to bass. “It is a very powerful diagnostic tool.”
Working with African primates, Goldberg’s group has employed these tools to identify dozens of new viruses, some of which might pose a threat to humans. With colleagues at the Wisconsin National Primate Research Center, he used the same techniques in 2015 to expose the culprit responsible for the death of Mahal, a beloved young orangutan at the Milwaukee County Zoo.
Mahal was a media star almost from the get-go. Born at a zoo in Colorado but rejected by his mother, the infant ape was ferried to Milwaukee in style aboard a private jet, and the Milwaukee Journal Sentinel routinely chronicled his early life. It was a shock in late 2012 when the five-year-old ape became ill and died unexpectedly.
The cause of Mahal’s death, Goldberg and his Milwaukee County Zoo colleagues discovered, was infection by an unrecognized species of tapeworm. Working only from a description of the orangutan’s clinical condition and small tissue samples, Goldberg and Primate Center colleague David O’Connor PhD’01 applied deep genetic sequencing tools. They found the unmistakable signature of a tapeworm that previously had been found only in weasels from Africa and Europe, deepening the mystery of how the ape became infected.
Tapeworms begin their life cycle as eggs in soil or water and transform into a larval stage in an intermediate host such as a rabbit or a mouse. When a predator or scavenger eats the rabbit or mouse, it consumes the tapeworm larvae, too, which develop into the adult form. If a tapeworm finds itself in an unfamiliar intermediate host, chaos can ensue.
“Mahal played the inadvertent role of intermediate host,” says Goldberg. “He must have eaten tapeworm eggs by accident, and the parasite found itself in an unusual place — an orangutan, instead of the mouse or rabbit it would typically infect. Without the proper checks and balances, it multiplied out of control and diffused throughout his body.”
But how did the ape become infected? Mahal was fond of eating dirt — something orangutans do naturally to aid digestion and obtain minerals. After a two-year investigation, it became apparent that Mahal was likely infected as an infant in Colorado, and that the tapeworm stayed quiet in his body for years.
Figuring out the story began with trapping wild animals at the Milwaukee County Zoo and sampling them for tapeworms. The zoo’s urban wildlife includes raccoons, skunks, coyotes, and mink. “We caught nothing but raccoons,” Goldberg says. “We kept catching the same ones.”
Ultimately, Goldberg found a genetic match in a tapeworm from an ermine in Colorado, with aid from a worldwide network of parasitologists and the U.S. Department of Agriculture’s National Parasite Collection. That finding supported the idea that Mahal was infected at the zoo where he was born. “This parasite is a new species,” says Goldberg. “You can’t classify these types of organisms using genetics alone. But if we someday are able to name the new species, I’d like to see it named after Mahal.”
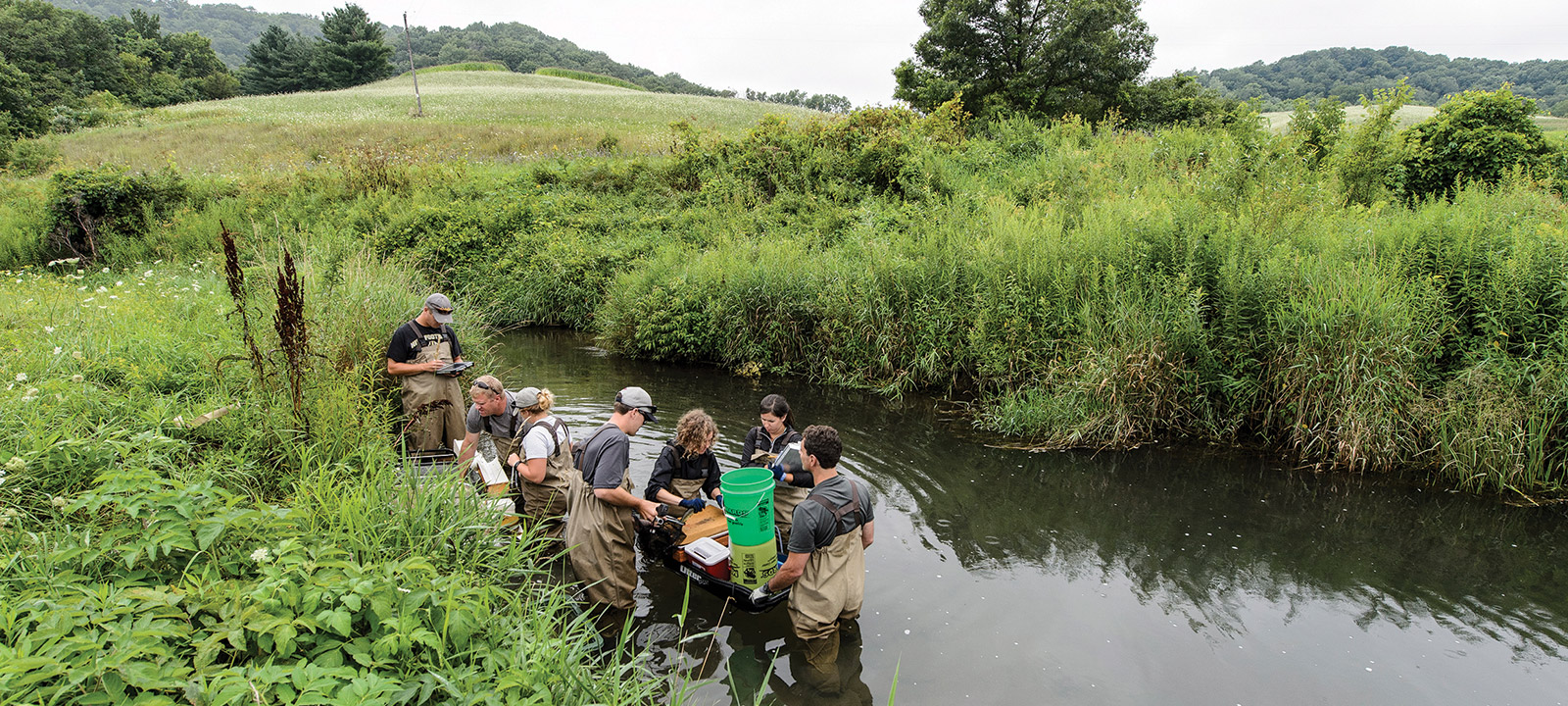
Hip deep in an ice-cold southwestern Wisconsin stream, wildlife biologist and veterinarian Bridget Baker ’01, MS’10, DVM’11 takes hold of a brown trout and expertly inserts a needle into its tail, probing for the largest vein. She carefully draws blood from the mottled, brassy brown fish, puts it in a labeled vial, and places it with the other samples she and her team have collected.
On this gunmetal gray July morning, Baker and her colleagues from the Wisconsin Department of Natural Resources are piggybacking with a fish- survey crew shocking 140-meter sections in two of the hundreds of streams that lace Wisconsin’s bucolic Driftless Area. The team works quickly to gather their samples of trout blood, 32 per stream. “We generally try to go with fish eight inches or longer,” Baker says. Fisheries biologists like Baker are on the front lines of confronting disease in Wisconsin’s fish. Today’s sampling exercise, they hope, will turn up no evidence of viral hemorrhagic septicemia, or VHS, a deadly infectious disease that arrived in the Midwest from Europe about a dozen years ago.
“We jokingly call it ‘Fishbola,’ ” says Goldberg of the virus that scientists speculate was introduced into the Great Lakes via the discharged ballast water of ships transiting the Saint Lawrence Seaway. The virus weakens blood vessels and causes severe hemorrhages in skin, muscle, and internal organs. It can survive in the water for at least 14 days, infecting some of the state’s most prized game fish, including largemouth bass, musky, pike, and trout. The virus has been found in Green Bay, Lake Superior, and some inland lakes, including Lake Winnebago. “We really don’t know how far it has spread,” Goldberg says.

Testing for “Fishbola,” the nickname for a deadly infectious disease that arrived in the Midwest from Europe a dozen years ago.
The fish blood samples will be subjected to an antibody test devised by Kathy Toohey-Kurth ’73, MS’84, a virologist with the Wisconsin Veterinary Diagnostic Laboratory and a UW clinical professor, in collaboration with Goldberg and funded by the UW Sea Grant Institute. Previously, testing for the virus required sacrificing the fish. “You had to take organs and grind them up to try and isolate the virus,” Goldberg says. “You don’t want to do that. So we developed a nonlethal test, which we are now applying to fish populations all around the state of Wisconsin.”
The key piece of information Goldberg and his collaborators want to provide to state wildlife officials is where the virus hasn’t shown up yet. “We really don’t want this virus spreading farther than it already has,” notes Goldberg, who, as a passionate fisherman, feels a personal stake in helping keep Wisconsin’s waters free of what could become an economic and quality-of-life disaster.
Despite the knowledge he and his colleagues have amassed about this deadly fish virus and similar diseases, Goldberg stresses there is more work to do.

Testing for the deadly virus VHS used to require killing the fish, before Goldberg and UW clinical professor Kathy Toohey-Kurth developed a nonlethal test.
“There are thousands if not millions of unknown viruses in the world. They all have little tricks up their sleeves that they have evolved to survive inside their hosts,” he says. “Although most will never emerge to cause us problems, some undoubtedly will.”
The pathogens that threaten us may best be thwarted by informed policy teamed up with the potent new tools deployed in the hunt for unknown diseases, but confronting them one at a time is like a game of Whac-a-Mole, Goldberg says.
Instead, he argues, we need to pay attention to larger forces in the world — environmental change, human encroachment on wildlife habitat, intensive agriculture, urbanization, changes in human population density — that may be “root drivers” of emerging diseases. The hypothetical bushmeat hunter who may have been the first to be infected with the chimpanzee precursor of the AIDS virus in Africa “may have lit the HIV match, but what stoked the fire was social change.”
Terry Devitt ’78, MA’85 is UW–Madison’s director of research communications. Bryce Richter is a photographer for University Communications and had to remove five ticks after this photo shoot.
Published in the Spring 2017 issue



Comments
No comments posted yet.