Genotopia
Eric Green ’81 has a vision of the future of medicine, and it looks a lot like William Gahl MD’76, PhD’81 and Louise Benge.
Green is the director of the National Human Genome Research Institute (NHGRI), an arm of the National Institutes of Health (NIH). This is more than alphabet soup — it’s the governmental organization that spent years sequencing the human genome. His vision is ambitious, but he doesn’t think it’s some utopia, and it’s not science fiction. “I want to be careful not to overpromise,” he says. “But I think in ten years, fifteen years, we’ll look back on some of our current medical treatments as being barbaric.” Green took charge of NHGRI in December 2009.
Gahl is the clinical director for NHGRI, making him responsible for the institute’s patient-based genomic research that goes on in NIH’s 234-bed hospital in Bethesda, Maryland. Gahl has also led — since its founding in 2008 — NIH’s Undiagnosed Diseases Program, a team of health professionals and researchers devoted to finding the causes of some of the most mysterious cases in modern medicine. It’s in this role that he met Benge.
Benge is not a doctor, nor a scientist of any kind. She’s a patient — best known in medical literature as “Family 1, Patient VI.1,” which is how she’s described in the February 3, 2011 issue of The New England Journal of Medicine. Benge does not know Green, but if his vision for the future of medical science is accurate, she will be among the first beneficiaries.
For thirty years, Benge has suffered excruciating pain, and were it not for Gahl, she still wouldn’t know why.
A native of Brodhead, Kentucky, Benge was in her early twenties when she first started feeling the effects of an ailment no one was able to define. When she walks for any distance at all — as little as a city block, perhaps, or a single flight of stairs — her legs become as hard as stone, and nearly as unresponsive. But they aren’t deadened and numb. They radiate searing pain.
For the first decade after this began, Benge tried to ignore the hurt until it went away. Eventually, she asked doctors — her primary-care physician and a host of specialists — for relief. They studied her, x-rayed her, and injected her with dye to trace the flow of her blood. And though they could see the visual evidence of her pain, they couldn’t determine what was causing it, let alone how to cure it.
Benge, they discerned, is the victim of her own circulatory system. A buildup of minerals causes her arteries to calcify and harden. This restricts blood flow, and though her body has tried to adapt by growing networks of tiny, collateral vessels, the blood that pumps to the cells in her legs and feet is still insufficient.
“My arteries are all calcified from the aorta on down,” Benge says. “It starts in the aorta and goes all down the lower extremities. [Doctors] can’t do surgery [to relieve the problem], because of the severity of it.”
Without knowing why Benge’s body was attacking her, her physicians could only rely on trial and error to search for relief. They put her on painkillers. They prescribed exercise to try to encourage increased blood flow. They even tried putting her on sodium thiosulfate, a medication used by patients in kidney failure. It can relieve the calcifying effects of dialysis, though it can also cause violent nausea. It did not help Benge. Nothing did.
“I was kind of at the end of my rope,” she says.
Eventually, her doctor, Karen Saylor, heard about Gahl and the Undiagnosed Diseases Program. Gahl was looking for patients at the end of their rope.
“The typical patient isn’t a good candidate for us,” Gahl says. “If we’re not your last resort, then coming here may be premature.”
Normally, it’s not rare for patients to get involved in an NIH-supported study. The institutes’ online list of clinical research projects, clinicaltrials.gov, names 916 current studies; of those, 138 are supported by NHGRI. But most of that research is what NIH calls extramural —essentially, NIH’s role is to help fund the studies through grants to labs at other institutions, such as the UW’s Genome Center of Wisconsin.
Gahl explains that only about 10 percent of NIH research is intramural, carried out in NIH government labs. And even among those studies, the Undiagnosed Diseases Program is special. Of the roughly 5,200 inquiries Gahl received in the program’s first three years, he accepted only 400. The program has to be exclusive because of the amount of effort expended on each case. Patients spend a week or more at the NIH hospital, meeting with specialists from virtually every field.
“It’s almost like, you name it,” Gahl says. “We have … neurologists and pediatric geneticists, but we consult with internists, pediatricians, dermatologists, rheumatologists, cardiologists, pulmonologists, and psychiatrists, physiometrists, rehabilitation medicine folks, pain and palliative care, people who are experts in bone disease, endocrinologists, gynecologists — any specialty, just about.”
Gahl was intrigued by the Benge case. Not only had she been through many different doctors, but the various elements of her disease were what he was looking for: “a new, unique, or remarkable physical finding or constellation of such findings, and maybe a clue that we can go after, an abnormal laboratory result.”
The clue in Benge’s case was that it wasn’t entirely unique. She could readily produce four other people with the same symptoms, and, importantly, two without.
Benge is the oldest of five children born to Clara and Bobby Proctor. Her siblings, in order of birth, are Derrell Proctor, Doug Proctor, Paula Allen, and Elizabeth Dodd, and all of them have the same calcified arteries — but their parents do not. Benge was the first to develop the symptoms of the mysterious disease, which was as strange to the Proctors as it was to their physicians.
“We used to kind of tease Louise for [occasionally] using a wheelchair,” says Allen. “Then, when I started feeling the same pain, I thought, well, I better have it checked into.”
One by one, the Proctor siblings all began showing signs of arterial calcification — sometimes first in the hands, manifesting like a form of severe arthritis, but always, eventually, in the legs and feet.
Allen says she can only walk for “ten to fifteen minutes at a slow speed, right now. My ankles and feet all hurt now. It’s not just my calves and legs. My feet hurt all the time. And they stay red. My feet, my legs, everything. I just ache.”
The presence of the symptoms in all five Proctor siblings but not in their parents indicated that the cause of the illness might be rooted in their genes — and that gave Gahl an idea for how to find the genetic culprit.
In May 2009, his team asked Benge and Allen to come to Bethesda.

Eric Green has been part of the human genome project “from day one” and now hopes to turn knowledge into treatments. Photo: Maggie Bartlett
To Green, Gahl isn’t just a doctor, and Benge isn’t just a patient. Together, they’re also elements in the effort to foster genomic medicine.
“What Bill is doing with the Undiagnosed Diseases Program is exactly what one has in mind as an incubator, taking the latest genomic technologies and applying them to figure out how best to diagnose diseases,” Green says. “[It shows] how to use genomics to do biomedical research and clinical research, and increasingly to change clinical practice.”
Genetics is the study of DNA and its role in heredity; genomics, however, is the study of a complete set of chromosomes — of all of a person’s genes — considering how those genes interact with each other and with the environment. Between 1990 and 2003, the human genome project was one of the highest profile efforts in biology, with the goal of mapping the entire human genetic code: the roughly 20,000 genes spelled out in some 3 billion nucleotides that constitute the reference set of human chromosomes. The effort cost some $3 billion, and laboratories in the United States, United Kingdom, Germany, France, China, Japan, and elsewhere took part.
Working with strands of DNA from a variety of people (but most of it from an anonymous donor from Buffalo, New York), teams of scientists began scanning, chromosome by chromosome, nucleotide by nucleotide — guanine, adenine, thymine, and cytosine — over and over, elucidating the chemical map of what it means to be human.
Green has been involved in the human genome project “literally from day one,” he says, working in labs, first at Washington University in St. Louis and then at NIH, sequencing and mapping, with a focus on chromosome 7. It was a massive effort, “biology’s equivalent to the moon shot,” Green says. But its aim was pure science, not directly connected to curing disease or helping patients.
“During the genome project,” he says, “we had a laser sharp, single goal: sequence the human genome. None of it was clinically oriented. It was just completely focused on a basic science endeavor.”
And that, Green believes, now has to change.
If sequencing the genome was biology’s moon shot, the discipline had its Neil Armstrong moment in 2003, when the complete sequence was published. In the eight years since, genomics has seen a surge in technology that enables scientists to sequence DNA with increasing speed and decreasing cost. But Green notes that NHGRI’s work can’t end with the publication of the reference human genome sequence. In his first year as director of NHGRI — and thus, at least administratively, America’s top genomicist — he led the organization to develop a new strategic plan. The institute must, he believes, find ways to apply genomics to solving the practical, clinical problems that real patients face. That plan, published in Nature in February 2011, proposes turning knowledge of the human genome into actionable medicine, diagnoses of diseases, and ultimately treatments.
“The bottom line,” he says, “is that … the future of genomics [is in] clinical applications.”
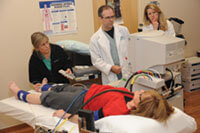
Louise Benge (on table) undergoes an arterial exam to evaluate circulation while NIH staffers Annette Stine, Kevin Smith, and Catherine Groden observe. Benge and her siblings went through a wide variety of tests while at the NIH facility in Bethesda, Maryland. Photo: Jeannine Mjoseth
Gahl’s Undiagnosed Diseases Program is one attempt to create those clinical applications and inject a spirit of practical discovery into genomics.
“A lot of people at NIH wanted to go back to the era of the 1950s and ’60s when new diseases were being diagnosed,” Gahl says. “So we thought it was an opportune time to start the program. So I hired a couple of people, and it was announced, and it was so popular with the press that NIH gave more money to it, and it just kind of grew from there.”
When Benge and Allen — and later Dodd and Derrell and Doug Proctor — went to Bethesda to meet with Gahl and his team, they found themselves subjects of a broad battery of examinations.
“We did every test imaginable,” says Benge. “Needles and testing and x-rays and CAT scans and blood work and more blood work and more blood work. PET scans and CT scans, ultrasounds. Just all kinds.”
Those tests revealed the extent of the damage that the illness had wrought. This was most visible in the large arteries in the Proctor siblings’ legs. In a healthy person, the femoral and popliteal arteries run like a highway down past the knee, carrying large amounts of blood to the lower leg and foot. Radiographs for Benge showed hers to be constricted and blocked — instead of the smooth flow of a highway, she had the crowded and crawling traffic of rush hour in a metropolis. Studies in cells indicated that the Proctors had low levels of an enzyme called CD73, and so Gahl’s team referred to the condition as ACDC, or Arterial Calcification due to a Deficiency of CD73.
But the scans, blood work, and x-rays only showed the effect of the disease. At the same time that Gahl’s team was running laboratory analyses on Benge and Allen, they were also looking for its origin in their genealogy.
The Proctors’ parents were third cousins. Though neither they nor any of the siblings’ ancestors showed any signs of the disease, the team theorized that they might have been passing it along nonetheless. What if several of them had been carriers of an ACDC mutation, a recessive gene that didn’t appear until Carla and Bobby Proctor married and brought two copies of that recessive mutation together?
“It was actually a pretty easy case, in a way,” Gahl says. Since the siblings’ parents were third cousins, “they share one–one hundred twenty-eighth of their genome sequence. Which means that slightly less than 1 percent of the entire genome is going to be homozygous. So we looked for a region in which all five individuals were homozygous and the parents were heterozygous. And there was only one region, and that region contained ninety-two genes. We were able to pick out which were the most likely candidates of those ninety-two and study them.”
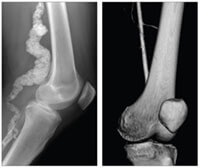
The femeropoplitial artery in Louise Benge’s leg (left) is heavily calcified (compare with a healthy artery at right).
The culprit, they found, was a gene called NT5E, which lives on chromosome 6 and governs production of CD73 enzyme. It had what Gahl calls a “nonsense mutation,” or defect. The discovery of that mutation — and of a similar mutation in four other patients (three sisters in northern Italy and a woman in California) who showed the same symptoms — led to publication in the New England Journal of Medicine and to the first announcement that the Undiagnosed Diseases Program had discovered a new disease.
“You ask the question, ‘Can you use new genomic technologies to better figure out the cause of rare diseases?’ Bill Gahl is answering that in spades,” Green says. “He’s launched this incredibly well-received program, which takes in patients from around the world, and for which the medical system has failed to deliver a diagnosis. … It’s a very clear demonstration that you can use these genomic techniques to figure out what is broken in patients with very rare genetic diseases.”
The Proctor siblings’ disease is, by any definition of the term, very rare. The federal government’s definition is found in the Orphan Drug Act of 1983, which classifies a disease as rare if it affects fewer than 200,000 Americans, or roughly one in every 1,500. The Proctor siblings’ particular form of arterial calcification affects only nine people on earth that we know of — or one in every 752,803,967 people.
But Green believes that while the Proctors’ disease may be very rare, the method of diagnosis might soon be nothing out of the ordinary — that the way Gahl worked to the root of the Proctor siblings’ problem also points medicine toward new ways to handle common medical situations. As an example, he says, consider warfarin, the blood thinner developed at UW–Madison by Karl Paul Link ’22, MS’23, PhD’25 in the 1940s and used to prevent strokes.
“Warfarin is a great drug,” Green says, “but people respond to it in wonky kinds of ways. It’s hard to get the dosage just right, because if you give people too much, they bleed out, and if you don’t give them enough, they clot. Getting that therapeutic window — you do it by trial and error. Very blunt and prone to problems.”
Recent genetic studies, he explains, have revealed the genetic variants that influence how a person metabolizes warfarin. Researchers are now conducting clinical trials to determine if using this genetic information about a patient improves a physician’s ability to find that therapeutic window and avoid the errors.
“We’ll see what the trials say,” Green notes, “but it is possible that they’ll demonstrate the effectiveness [of genetic testing]. If that happens, then it will become the standard of care — that before you give somebody warfarin, you quickly give them a DNA test and figure out what kind of dosing you should use.”
In Green’s vision, genome sequencing would become as standard to medicine as physicals and blood tests. Children might have their genomes sequenced at birth, the resulting information then helping physicians to understand what kinds of illnesses they might be subject to, and which kinds of medicines might be most effective, and which potentially dangerous. Doctors might learn which individuals were at risk for what kinds of cancers, and how those cancers are best treated.
“Maybe ten years, maybe fifteen years from now, we wouldn’t do genetic tests on patients in real time,” he says. “Rather, maybe the patient would already have had his or her genome sequenced, and that information would be part of the electronic medical record.”
But ten or fifteen years is a long time to wait for Louise Benge and her siblings. For them, the promise of the future is less important than the pain of the present.

Bill Gahl (right) heads the NHGRI’s Undiagnosed Diseases Program, which seeks out the causes of mysterious ailments. Here, he’s shown with (from right) Louise Benge, Paula Allen, and Manfred Boehm, a researcher at the Center for Molecular Medicine. Photo: Jeannine Mjoseth.
Though Green has high hopes for genomics to change the practice of medicine, he also knows that there’s much work to be done to translate genomic knowledge into cures.
“What Bill Gahl’s doing is as much a research endeavor as anything else,” he says. “Just because you can diagnose a disease doesn’t mean you can treat it.”
And Gahl is acutely aware of this distinction — and that treatment is not in his charter. It is, rather, a personal desire.
“We don’t have a goal of treating patients,” he says. “We just want to do it when we see the patients, and they have no alternative.”
And so after the Undiagnosed Diseases Program found the cause of the Proctor siblings’ disease, its members also began working on a regimen to bring the Proctors relief. Having determined the base of their problem — that their genes won’t produce the enzyme that would keep their arteries from calcifying — Gahl and his team have worked up an experimental treatment that would obviate the need for the missing enzyme.
But the path to relief isn’t simple, and the standards of science, and of medical regulation, don’t make Gahl’s job any easier.
“In general, the Food and Drug Administration applies the same standards for rare diseases as for common diseases,” Gahl says. “If you have a one-off disease … you still have to do pre-clinical animal toxicity studies. And nobody’s going to do that,” he says — no one’s going to spend the time and energy to develop a drug with a market of only nine people in the world.
So since their journey to NIH, the Proctors have been on essentially the same treatment plan they had before: exercise to increase blood flow. Each day, Benge and her siblings hit their treadmills to walk for as long as they can stand, then rest, then start walking again.
“It was wonderful to finally find out what was causing our problems,” Benge says. “And they think they found some medication that can help us, but they have to get it approved. If they can get that done, get us started on the rounds of medicine and stuff, it would be even more wonderful, if it helps us.”
John Allen is senior editor of On Wisconsin Magazine.
Published in the Fall 2011 issue



Comments
douglas proctor October 16, 2011
I would like to thank Dr. Gahl and all the doctors,researchers, and all involved for the time amd effort that has gone into finding all this out.To Dr. Saylor for sending all of us to the N.I.H. .It blows most of the peoples minds that i have talked to that the name came up as ACDC,,, most think that i am kinding them until i explain it.Reason they know that I listened to the band ACDC for years,,For all the tests that you-all have run thank you again and I know that we all appreciate it.
Dan Vera May 5, 2012
What a super story and diagnostic road map. One wishes you Godspeed in making medicine a ladder of understanding. Adapting your thought process to an algorithm should indeed be a priority. May your virtue guide you further that the creative nature of your work blossom into a flower of curative effects.
I got disbarred for suing a medical doctor for lying to a patient and harvesting her uterus under the pretext of her having a 3rd degree prolapse. I was told I had filed a frivolous lawsuit. After I showed the judge the consent form with changes made with white-out; a bi-lateral tubal ligation became a hysterectomy by mystery, and yet he would not allow the case to go to trial.
Doctors like that make doctors like you look like fools. And I know you are not fools. Please tell the governor that gomez v matey (TX) was not a frivolous lawsuit.
Dr David Brabon February 21, 2013
I am so pleased to work in the same community as Dr Karen Saylor. She is one of the most brilliant physicians I know. She did a magnificent initial work up and recognised a unique illness. She didn’t brush it off as some would, but pushed through to a clear diagnosis. This also meant that she is so well verse in the general scope of medicine,syndromes and disease processes that when something this unusual occurs she can readily recognize it and refer it appropriately. At times I fear we don’t value the gifted in our midst!